Finding Antibiotics in Unexpected Places - Medicine from Mammoths?
Finding Antibiotics in Unexpected Places - Medicine from Mammoths?
By Jakob Jung
Microbes are everywhere. They live in soil, seas, rocks, and waters, and have been colonizing our planet for billions of years before humans appeared. Some even survive boiling volcanic springs above 100 °C, others endure glacial ice below -20 °C. They also live inside us. The human microbiome—the vast community of microorganisms in and on our bodies—works in harmony with our immune system to protect us from disease. But when certain microbes invade places they don’t belong to, like our airways, bloodstream, or wounds, they can become dangerous. Pathogens such as Mycobacterium tuberculosis (which causes tuberculosis) and Yersinia pestis (which causes plague) cause infections that, for most of human history, were untreatable and often fatal. Without effective drugs, these microbes can be very deadly claiming the lives of hundreds of millions of people throughout history.
The Discovery That Changed Medicine
That changed in 1928, when Alexander Fleming noticed something remarkable: bacteria were dying in places where a specific mold was growing. The fungus Penicillium notatum (later replaced in production by Penicillium chrysogenum ) produced a substance that killed microbes. That substance became penicillin—the first widely used antibiotic. It sparked a revolution in medicine, transforming once-deadly infections into treatable conditions and saving numerous human lives ever since.
Nature’s Arsenal of Antibiotics
Penicillium fungi produce penicillin to kill bacteria, and thereby gain a competitive advantage in accessing nutrients. Scientists later realized that nature was full of such microbial battles. Bacteria, fungi, and other microbes possess a diverse arsenal of antimicrobial compounds to kill other microbes. This insight drove the “golden age of antibiotic discovery” from the 1940s to 1970s, when most of the antibiotics we rely on today were found by investigating chemical compounds from these organisms.
Unfortunately, bacteria can develop resistance to these antibiotics, rendering them ineffective and creating a growing global health crisis called antimicrobial resistance (AMR), which is responsible for millions of deaths worldwide. Therefore, governments and scientists need to identify new antimicrobial compounds, using new methods.
Beyond Microbes: Unexpected Sources of Antimicrobials
Microbes aren’t the only source of antimicrobial weapons. Plants, deep-sea sponges, and even our own immune system can produce molecules that kill bacteria. Human cells, for example, produce antimicrobial peptides—tiny proteins that punch holes in microbial invaders. These antimicrobial peptides share common structures and sequences; therefore, they can be predicted bioinformatically purely from DNA genome sequences. With the rapid advances in DNA sequencing and machine learning, researchers can quickly search across multiple genomes for hidden antimicrobial peptides. These digital tools scan vast databases of microbial and animal DNA to predict which sequences may encode new antimicrobial peptide-based drugs.
Mining DNA for New Medicines
And here’s the twist: DNA doesn’t just come from living creatures. Ancient DNA from extinct animals can also be sequenced and is available in databases. Recently, scientists explored the genomes of long-gone species—including the woolly mammoth—and uncovered several promising antimicrobial peptides. One mammoth-derived peptide, called mammuthusin-2, even effectively inhibited the growth of pathogenic bacteria in mouse wounds. This approach, sometimes called “molecular de-extinction,” hints at a radical idea: life forms from the past could help us fight one of today’s greatest threats—antimicrobial resistance (AMR).
The search for new antibiotics is a race against time. By looking not only to living organisms but also to the genetic echoes of the past, researchers are expanding the toolbox of modern medicine in surprising ways. From molds to mammoths, nature may still hold the keys to our survival.
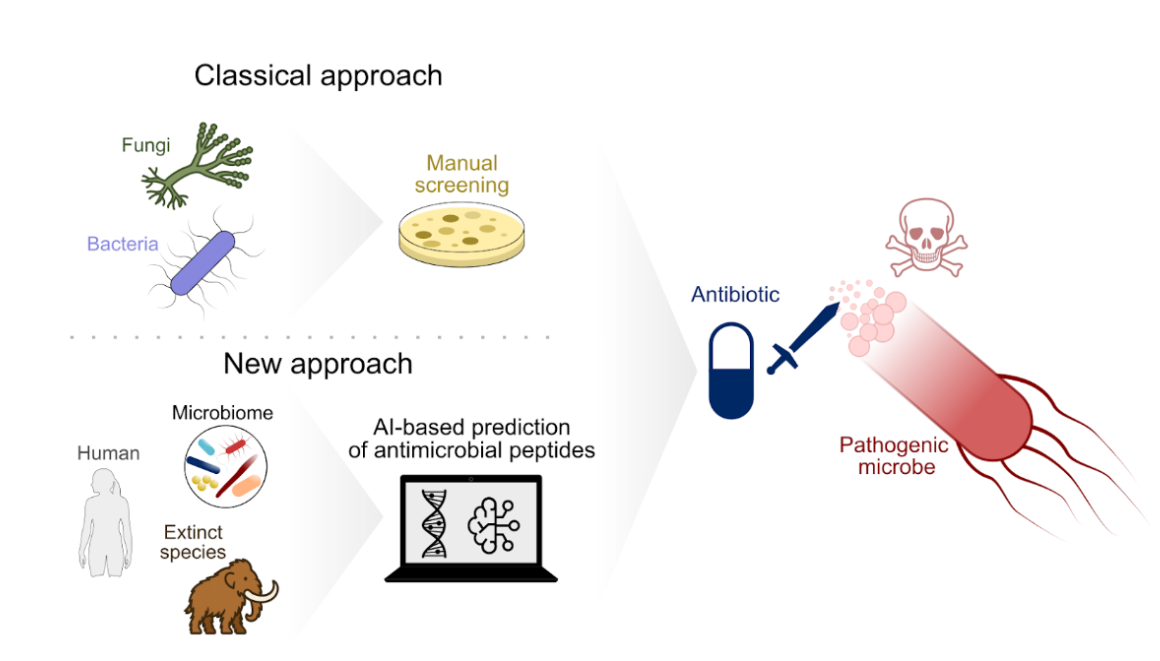
Antibiotics were initially discovered in fungi and soil bacteria, by manual experimental screens of these microbes. But also other life forms, including bacteria, humans, and even extinct mammals such as mammoths, have evolved to produce antimicrobial peptides capable of killing pathogenic microbes. These can be identified using artificial intelligence-based computational tools that search big genome databases for antimicrobials.
About the author:
Jakob Jung is a doctoral researcher at the Institute of Molecular Infection Biology (IMIB) in Würzburg, Germany. His work focuses on the intersection of antibiotics, the microbiome, and human health, with a particular interest in health-related bioinformatics and innovative treatment strategies.
He first developed a passion for microbiology during his bachelor’s studies, where he combined theoretical training with hands-on laboratory experience. This led him to pursue a master’s degree in Cellular Biotechnology, specializing in computational biology, metabolic modeling, and next-generation sequencing (NGS) techniques.
Since 2020, Jakob has been exploring how computational tools can deepen our understanding of microbial interactions and support the development of new therapeutic approaches.